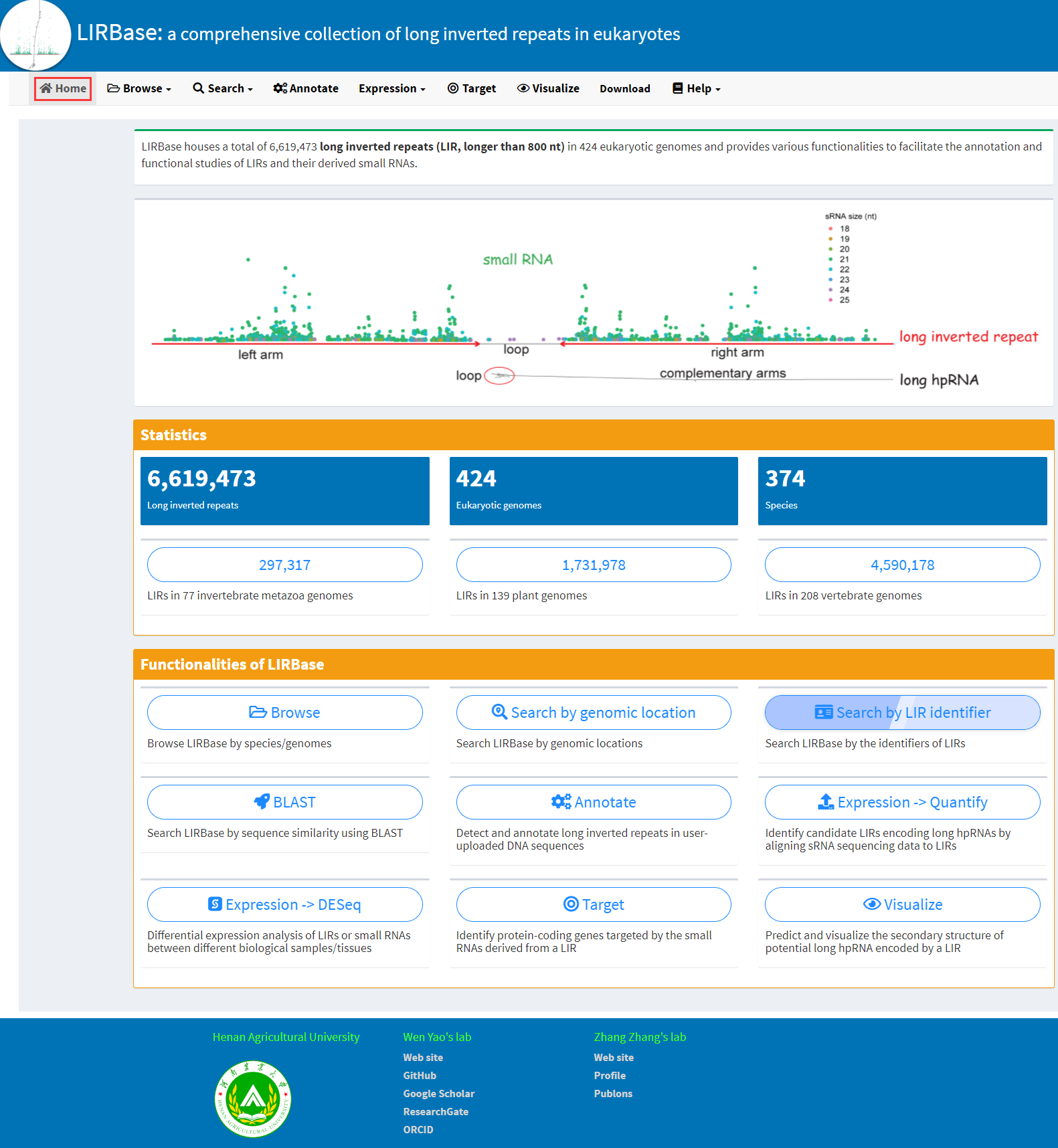
MiteFinderII: a novel tool to identify miniature inverted-repeat transposable elements hidden in eukaryotic genomes | BMC Medical Genomics | Full Text
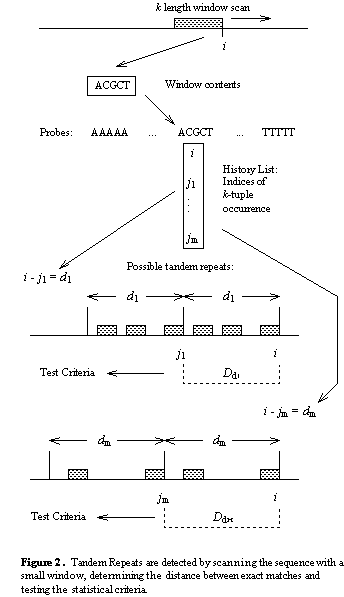
![PDF] detectIR: A Novel Program for Detecting Perfect and Imperfect Inverted Repeats Using Complex Numbers and Vector Calculation | Semantic Scholar PDF] detectIR: A Novel Program for Detecting Perfect and Imperfect Inverted Repeats Using Complex Numbers and Vector Calculation | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/a20f6bea6d62205d58f8bcd56fa3411c859aad47/3-Figure1-1.png)
PDF] detectIR: A Novel Program for Detecting Perfect and Imperfect Inverted Repeats Using Complex Numbers and Vector Calculation | Semantic Scholar

The distribution of inverted repeat sequences in the Saccharomyces cerevisiae genome | Current Genetics
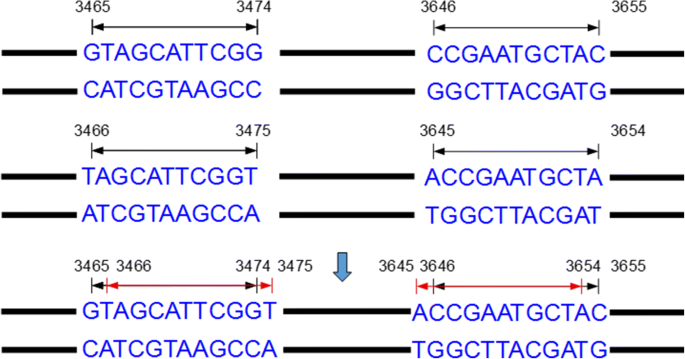
MiteFinderII: a novel tool to identify miniature inverted-repeat transposable elements hidden in eukaryotic genomes | BMC Medical Genomics | Full Text
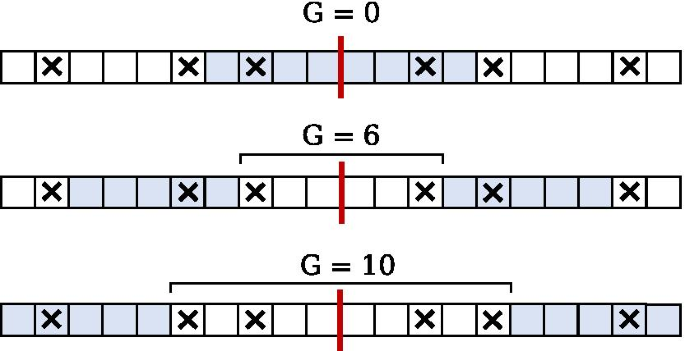
IUPACpal: efficient identification of inverted repeats in IUPAC-encoded DNA sequences | BMC Bioinformatics | Full Text
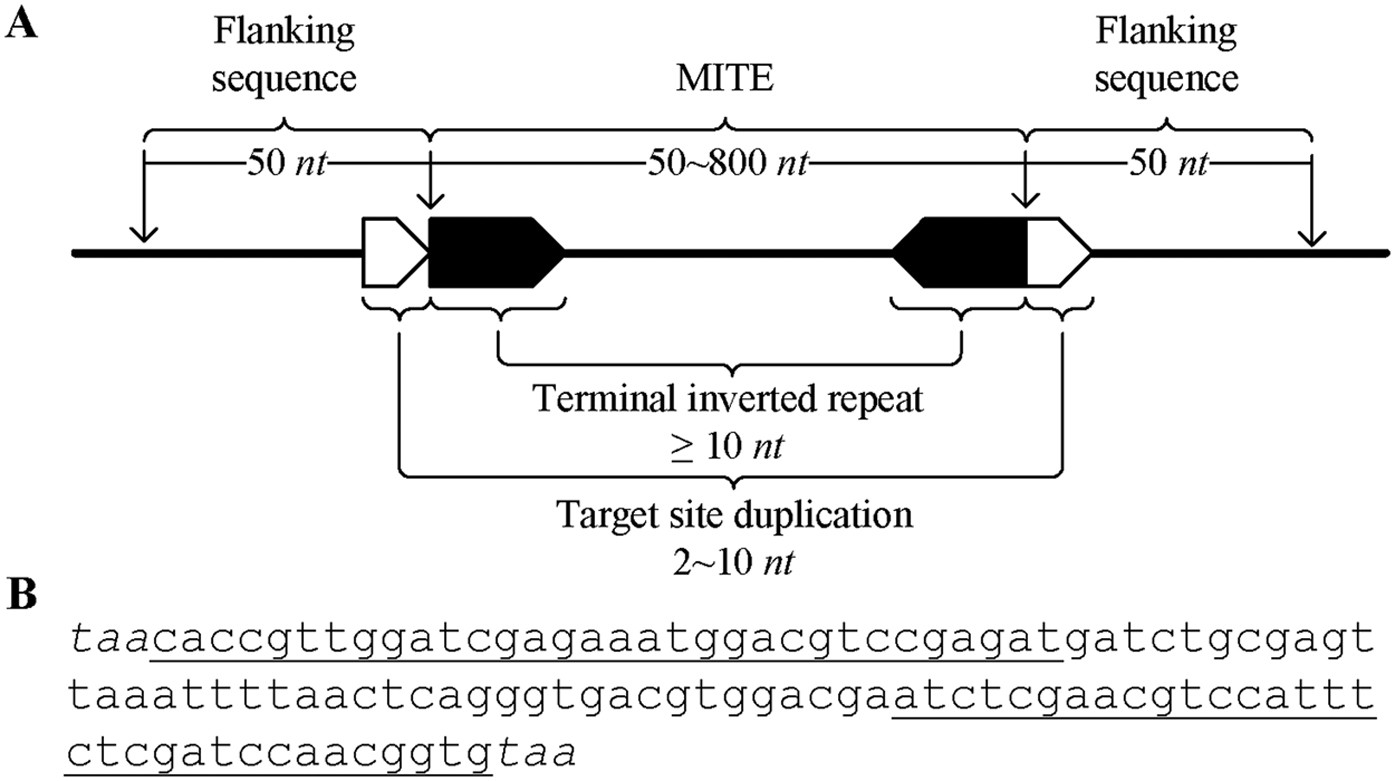
detectMITE: A novel approach to detect miniature inverted repeat transposable elements in genomes | Scientific Reports

The chloroplast genome of the lichen‐symbiont microalga Trebouxia sp. Tr9 (Trebouxiophyceae, Chlorophyta) shows short inverted repeats with a single gene and loss of the rps4 gene, which is encoded by the nucleus -

Expanded inverted repeat region with large scale inversion in the first complete plastid genome sequence of Plantago ovata | Scientific Reports
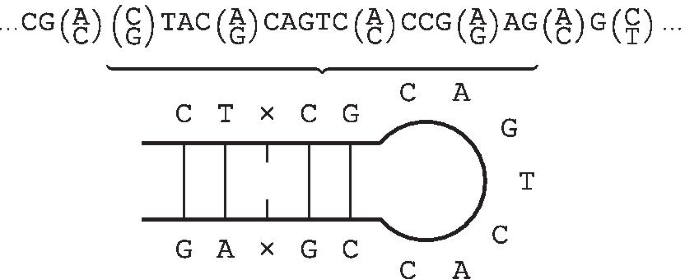
IUPACpal: efficient identification of inverted repeats in IUPAC-encoded DNA sequences | BMC Bioinformatics | Full Text

npInv: accurate detection and genotyping of inversions using long read sub-alignment | BMC Bioinformatics | Full Text